buccalfatremoval
I love cigarettes
- Joined
- Jun 30, 2024
- Posts
- 117
- Reputation
- 81
Alright boyos, I’m sure you all know what HGH is, but today I want to dive deep into it.
Think of it as a research paper about HGH with a little theory at the end.
This is going to be a long read.. maybe.
1.INTRODUCTION
Human growth and development are tightly regulated by steroid and peptide hormones. Steroid hormones, including androgens (e.g., testosterone) and estrogens (e.g., estradiol), play a critical role during puberty by driving the pubertal growth spurt, promoting bone maturation, and inducing epiphyseal closure, which ultimately ends linear growth. They are also responsible for the development of secondary sexual characteristics. Genetic defects in enzymes involved in steroid hormone synthesis can result in growth and developmental disorders [1–5].
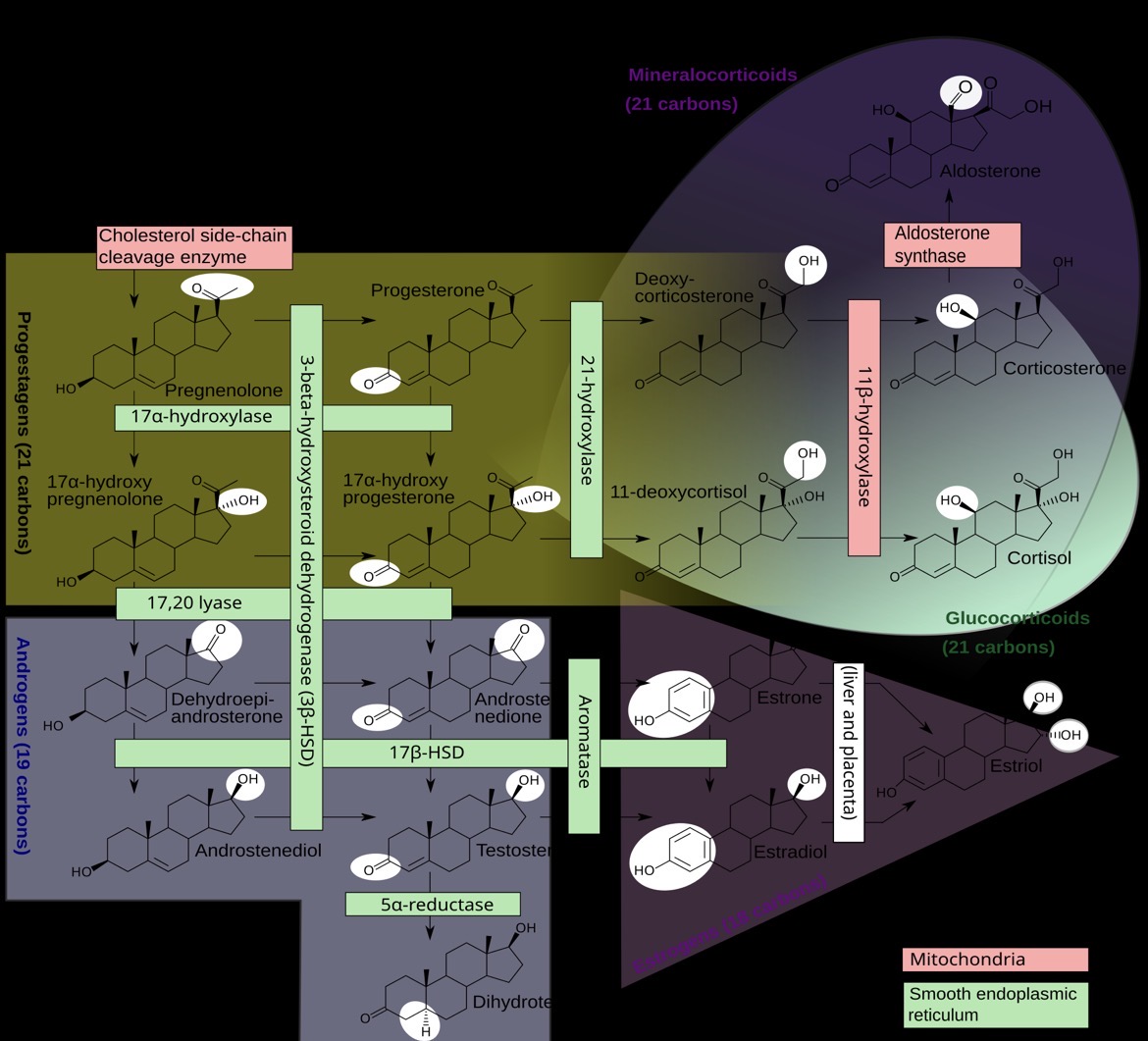
(steroidogenesis pathway highlighting key enzymes, genetic defects (e.g., 21-hydroxylase deficiency) disrupt hormone synthesis and can lead to growth and developmental disorders.)
Growth hormone (GH), a peptide hormone produced by the anterior pituitary gland, is a key regulator of growth and metabolism, particularly during childhood and adolescence [6]. GH exerts direct effects on tissues such as bone and cartilage and indirect effects by stimulating the production of insulin-like growth factor-1 (IGF-1), primarily in the liver. IGF-1 further promotes somatic growth. GH secretion is regulated by growth hormone-releasing hormone (GHRH) from the hypothalamus, which binds to GHRH receptors (GHRHR) on pituitary somatotroph cells. GH then acts through growth hormone receptors (GHR) expressed on target tissues. Steroid hormones modulate this GH–IGF-1 axis by influencing both GH secretion and IGF-1 production, highlighting the complexity of hormonal control in human growth.
The human growth hormone gene (GH1) is located on chromosome 17 (region 17q22–24) and is part of a cluster of five related genes, including GH2 and chorionic somatomammotropin genes. This cluster spans about 65 kb and reflects shared evolutionary origins and coordinated regulation. GH2 is specifically expressed in the placenta, suggesting a role in pregnancy. The mature human growth hormone (hGH) protein consists of 191 amino acids, formed after cleavage of a signal peptide from a larger precursor. Structurally, hGH has a four-helix bundle stabilized by disulfide bonds, which is essential for binding to the growth hormone receptor (GHR). Its amino acid sequence is highly conserved, especially in receptor-binding regions.
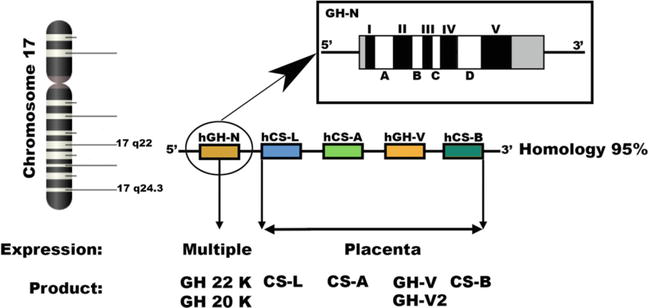
(Organization of the GH1 gene cluster on chromosome 17 showing related growth hormone and placental hormone genes.)
GH is synthesized and secreted by somatotroph cells in the anterior pituitary gland under hypothalamic control. Growth hormone-releasing hormone (GHRH) stimulates GH release, while somatostatin inhibits it; ghrelin also promotes GH secretion. GH is released in a pulsatile manner, with peak secretion during slow-wave sleep. Its secretion is influenced by multiple factors, including circadian rhythm, nutrition, stress, exercise, age, and body composition. In circulation, GH is mainly bound to growth hormone-binding protein (GHBP), which regulates its availability and activity and participates in feedback control.
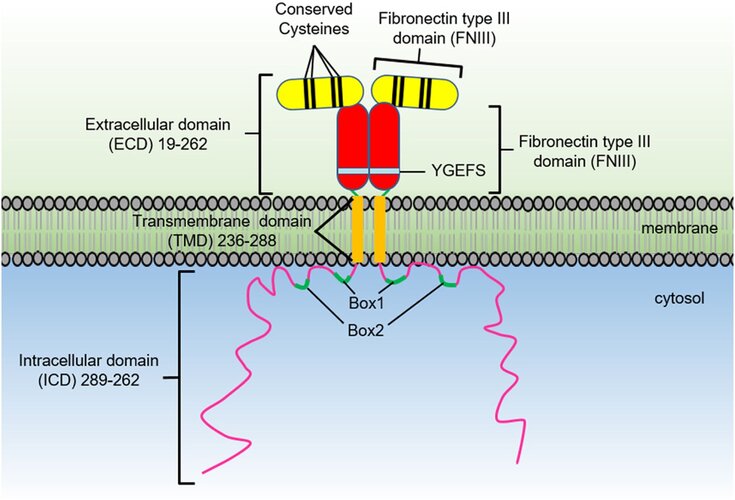
GH has widespread physiological effects beyond linear growth. It promotes bone elongation, muscle development, protein synthesis, lipid metabolism, immune function, and maintenance of bone density. Its actions are mediated through binding to the GHR, a cytokine receptor expressed in many tissues such as liver, bone, and muscle. This interaction activates intracellular signaling pathways, primarily the JAK2/STAT5 pathway, leading to gene transcription.
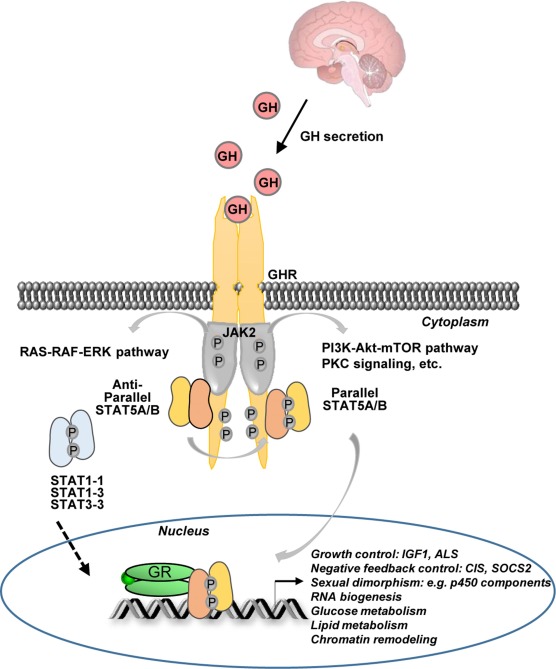
(Growth hormone receptor activation and downstream JAK2/STAT5 signaling pathway.)
A major downstream effect of GH signaling is the production of insulin-like growth factor-1 (IGF-1), mainly in the liver. IGF-1 stimulates tissue growth and also exerts negative feedback on GH secretion. While many growth-promoting effects of GH are mediated by IGF-1, GH also has direct actions, including stimulation of bone cells, regulation of glucose metabolism (often opposing insulin), and promotion of fat breakdown.
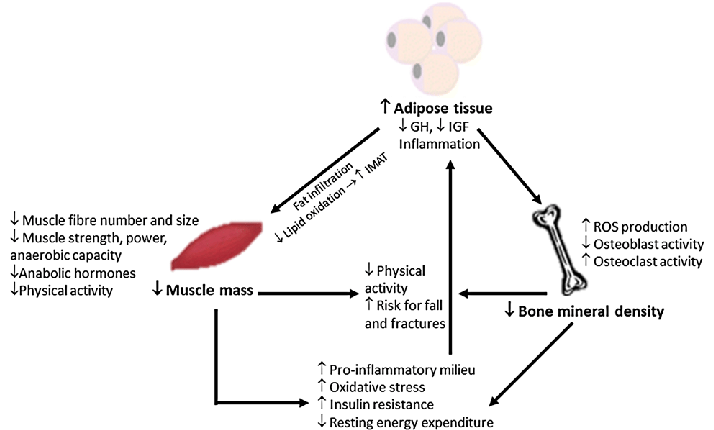
(Physiological effects of growth hormone on skeletal growth, muscle development, and metabolism.)
Disruptions in the GH pathway—often due to mutations in GH1 or related genes—can result in growth disorders. Isolated growth hormone deficiency (IGHD) leads to reduced GH production and short stature, while mutations in the GHR cause GH insensitivity (e.g., Laron syndrome). These mutations may impair GH secretion, receptor binding, or downstream signaling.
This study focuses on understanding how GH structure, evolutionary conservation, and disease-causing mutations affect its interaction with the GHR, aiming to clarify the molecular basis of GH-related disorders.
2. AMINO ACID SEQUENCE
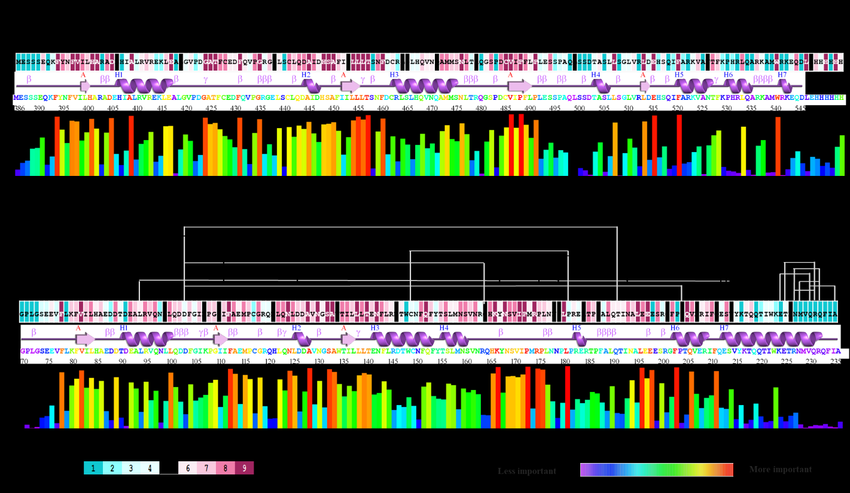
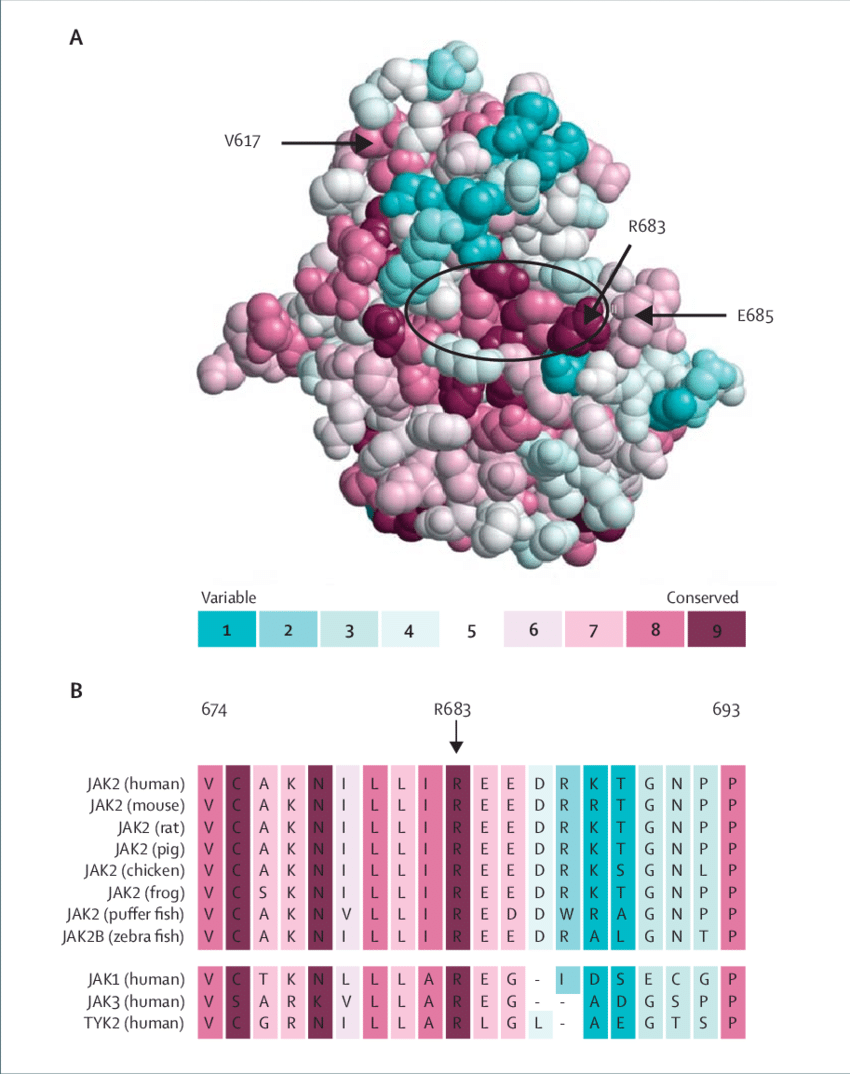
(Evolutionary conservation analysis of GH residues mapped onto protein structure using ConSurf scoring.)
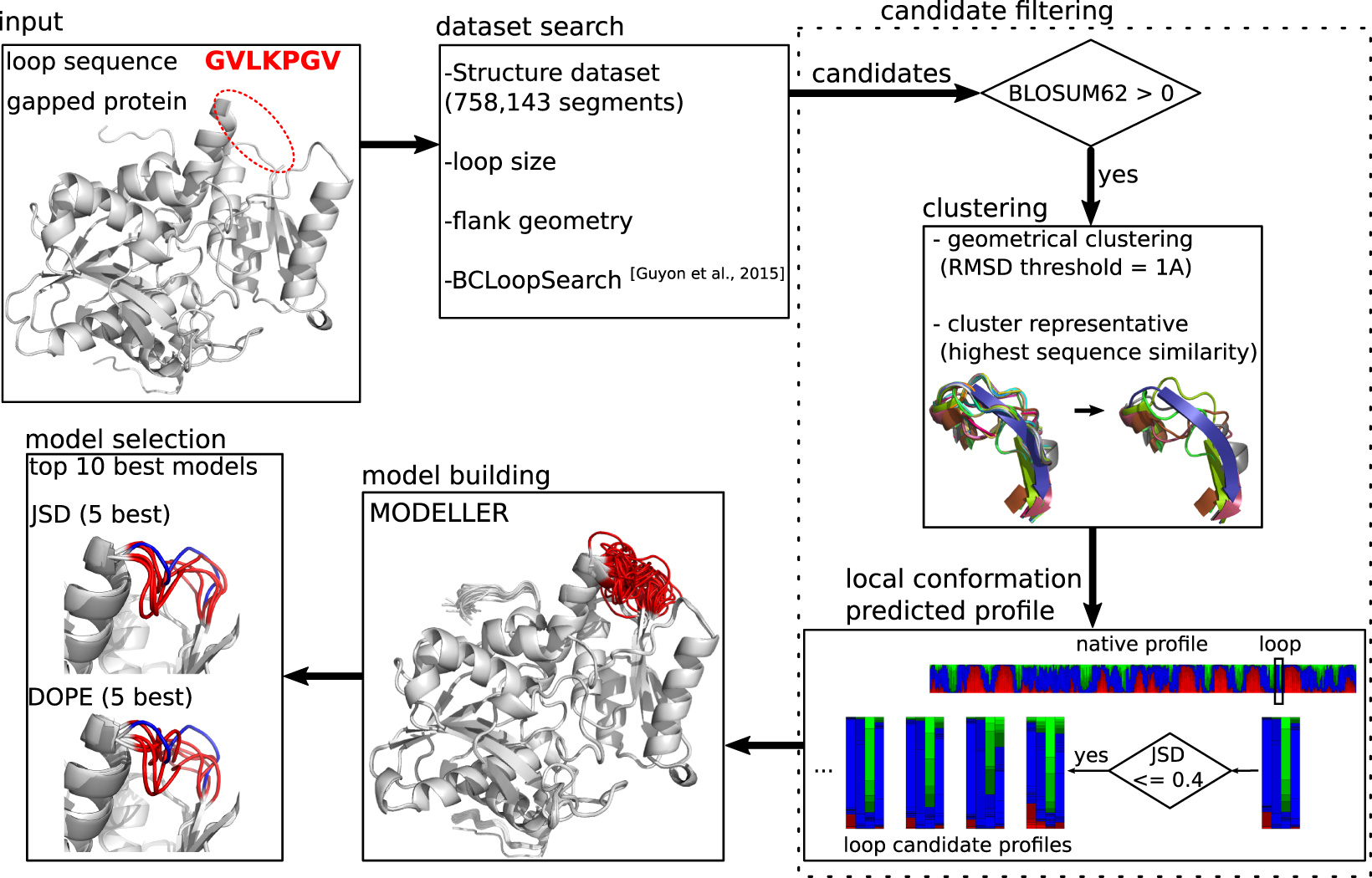
(Protein loop modeling of missing residues in GH structure using computational structure refinement methods.)
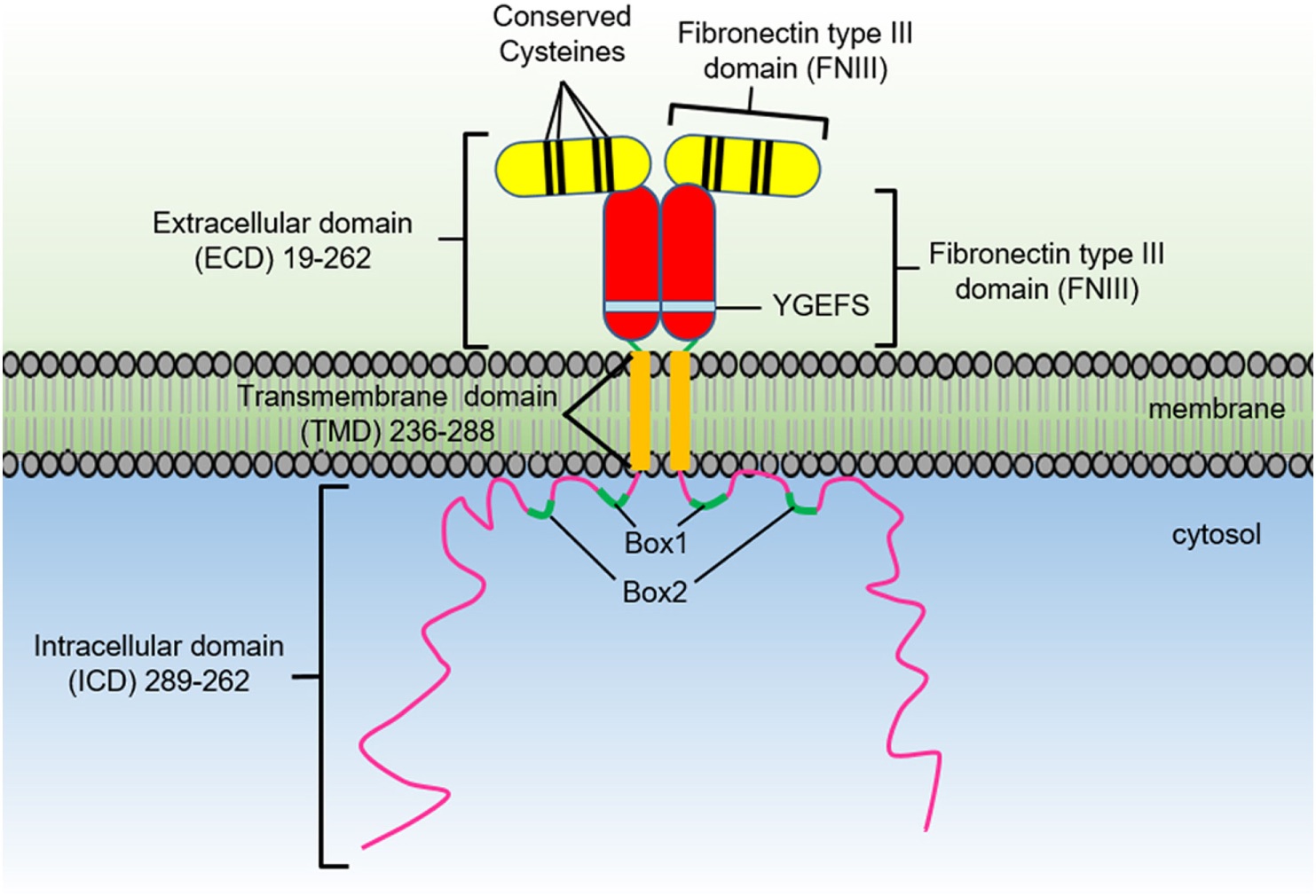
(Human growth hormone bound to its receptor showing key interaction sites involved in binding and signaling.)
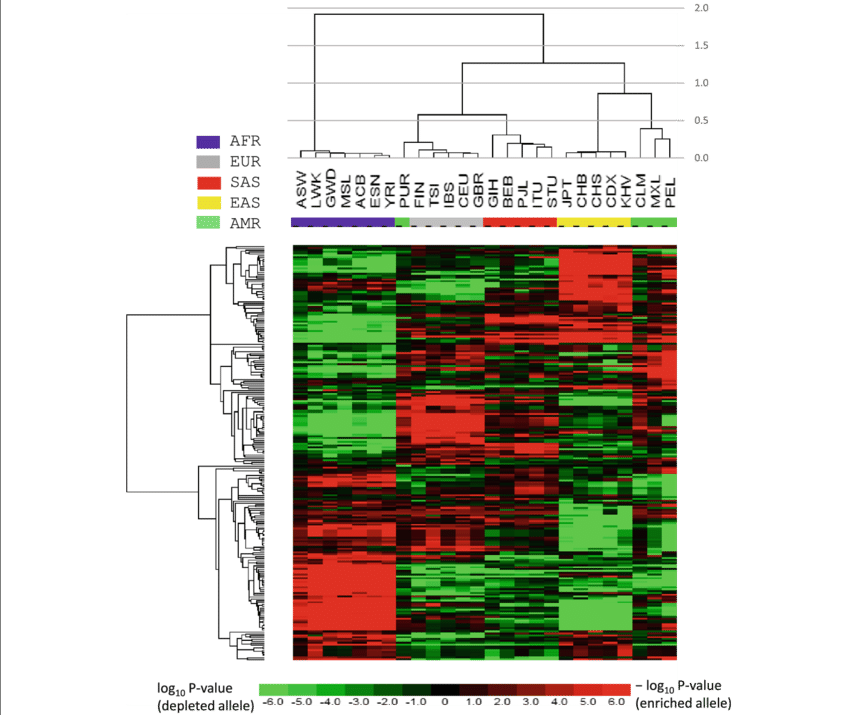
(Population-level allele frequency distribution of GH population from large-scale genomic sequencing databases.)
Birds show lower conservation, with 55–57% identity and 72–74% similarity, as seen in turkey, chicken, and ostrich. The lowest conservation is found in fish species, where identity drops to 36–44% and similarity to 52–61%, with Atlantic salmon showing the greatest divergence. Overall, GH is highly conserved in mammals but becomes increasingly divergent in birds and especially fish, reflecting evolutionary distance while maintaining essential functional regions.
Table 1.
Sequence comparison of human growth hormone with growth hormone sequences from other species. The table shows the NCBI and UniProt sequence identifiers, UniProt sequence name, sequence length, percentage of sequence identity, percentage of sequence similarity, and signal peptide length for each species compared to human GH.
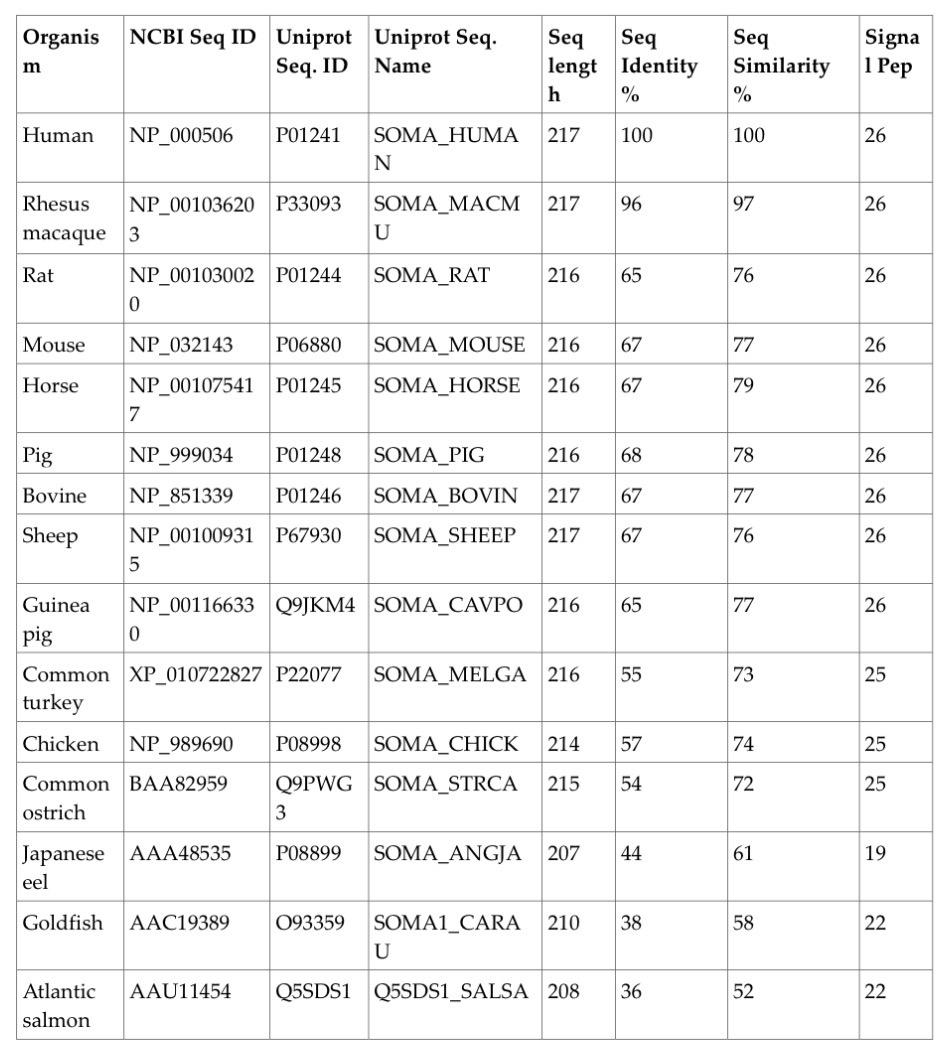
The length of the mature growth hormone protein is highly conserved across the mammalian species, ranging from 216 to 217 amino acids, similar to the 217 amino acids in human hGH. The signal peptide length is also conserved at 26 amino acids in most mammals, with a slightly shorter signal peptide of 25 amino acids in the avian species and 19-22 amino acids in the fish species.
3.1.2 Analysis of human GH mutations and their locations in GH protein
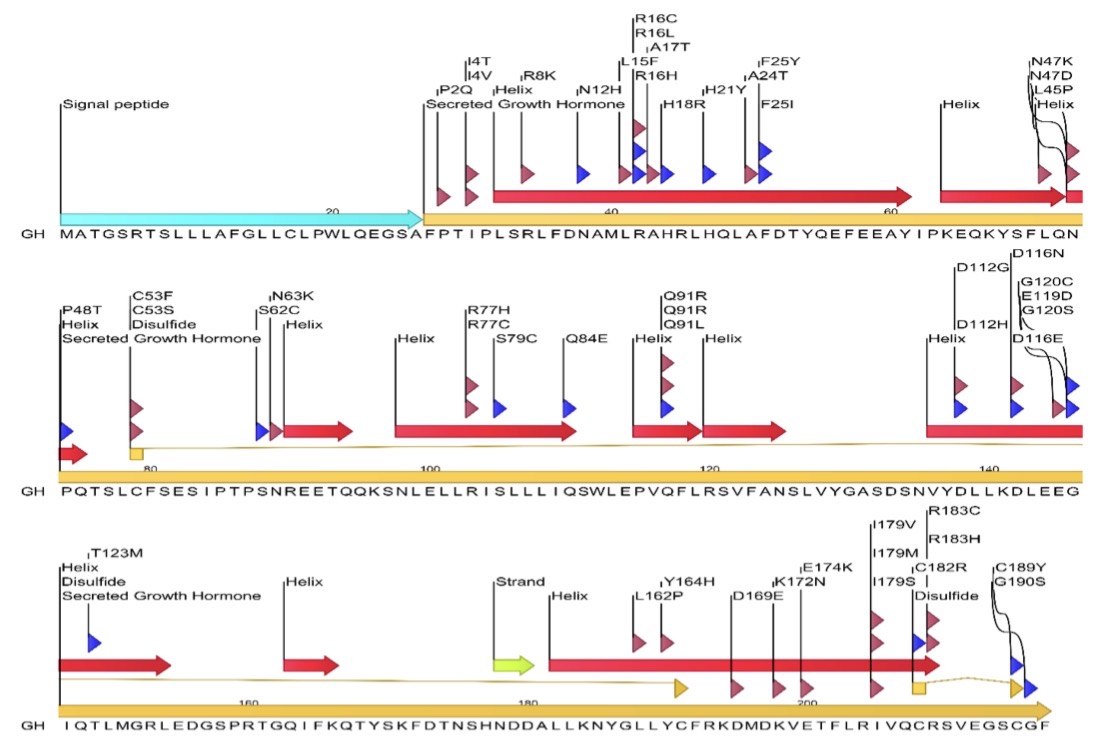
Figure 1 illustrates the linear structure of the human growth hormone (GH) protein and the distribution of reported mutations associated with disorders such as growth hormone deficiency (GHD) and Laron syndrome (LS). The GH protein contains four main α-helices (A–D), two disulfide bonds, and connecting loop regions, all of which contribute to its structural stability and receptor binding.
Many disease-associated point mutations are spread throughout the protein, but a high concentration occurs in the α-helical regions, especially helix A (residues 9–34) and helix D (residues 158–190), which are critical for growth hormone receptor (GHR) interaction. Additional mutations are located in loop regions between helices, including the N-terminal region and loops connecting helices A–B, B–C, and C–D. These mutations affect amino acids involved in structural stability and receptor binding, potentially disrupting GH function and leading to impaired signaling. Overall, the figure highlights that both helical and loop regions contain functionally important mutation hotspots linked to GH-related disorders.
TABLE 2.
Table 2 compiles reported disease-associated mutations in the human growth hormone (GH) protein, detailing amino acid substitutions, their sequence positions, and linked clinical outcomes such as growth hormone deficiency (GHD) and related syndromes. It also includes information on the functional impact of each mutation, such as reduced receptor binding, impaired secretion, or altered signaling activity.
In addition, the table provides dbSNP identifiers (where available) and references to the original studies reporting each mutation. Sequence positions are noted in both the full-length precursor protein and the mature GH form, accounting for the presence or absence of the 26-amino acid signal peptide. Overall, the table highlights how specific point mutations across the GH sequence contribute to functional disruption and disease development.
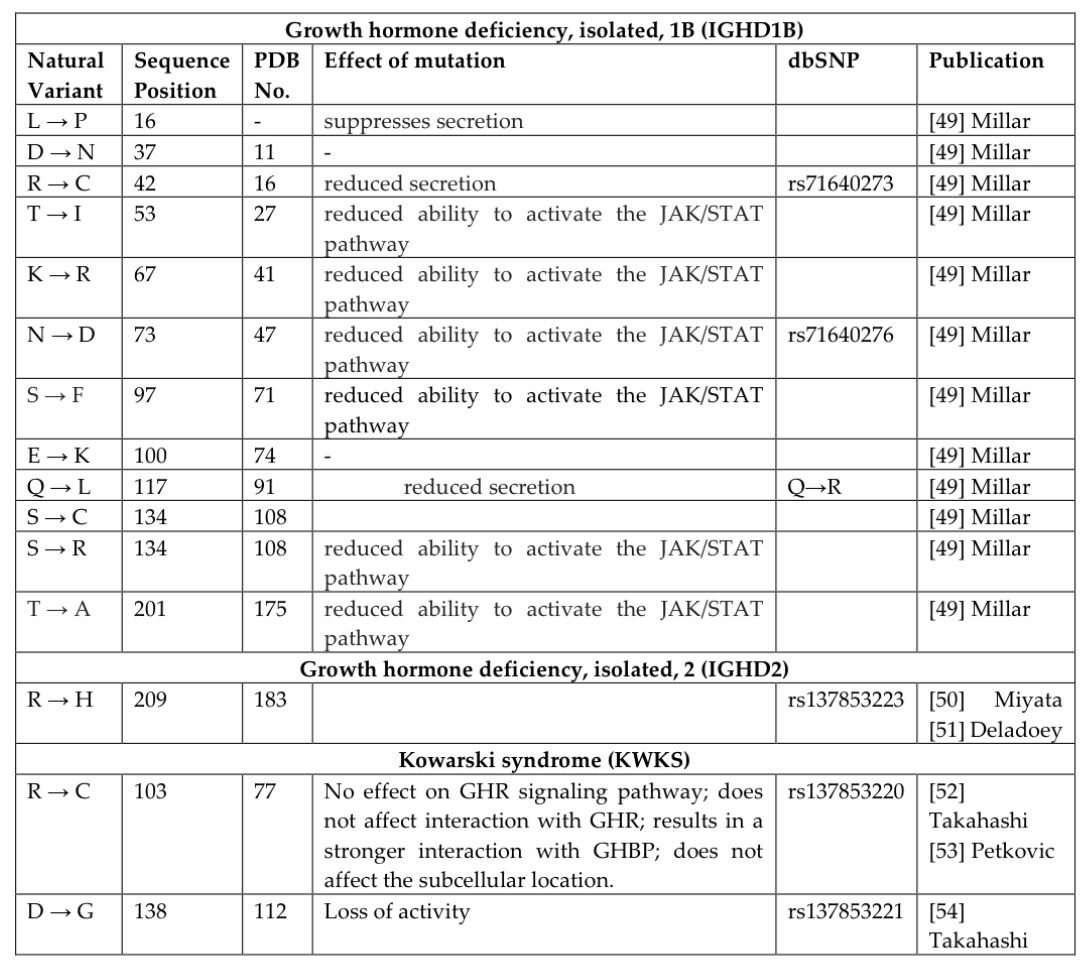
Disease-causing mutations in human growth hormone (GH), retrieved from the UniProtKB database, are mainly associated with isolated growth hormone deficiency (GHD) and Kowarski syndrome (KWKS). GHD is typically an autosomal recessive condition characterized by short stature, low but detectable GH levels, delayed bone age, and responsiveness to GH therapy. In contrast, Kowarski syndrome is defined by bio-inactive GH, resulting in normal or slightly elevated GH levels, very low IGF-1, and improved growth with GH replacement therapy.
These mutations are distributed throughout the GH protein and affect its function in different ways. Some mutations, such as L16P and Q117L, mainly impair GH secretion. Others, including T53I, K67R, N73D, S97F, and T201A, disrupt activation of the JAK/STAT signaling pathway, reducing biological activity. The R209H mutation is linked to autosomal dominant IGHD type 2 (IGHD2). Specific variants such as Thr3, Arg16, Asn47, Gln91, Arg183, Arg77, and Asp112 are associated with GHD and KWKS.
Certain mutations show distinct functional effects: R103C (KWKS) does not disrupt receptor signaling but increases binding to growth hormone-binding protein (GHBP), while D138G leads to a complete loss of biological activity. Overall, these mutations highlight how changes in different regions of GH can lead to diverse clinical outcomes through effects on secretion, receptor interaction, and intracellular signaling.

Figure 2: A multiple sequence alignment of human growth hormone alongside growth hormone sequences from various vertebrate species. The alignment encompasses the N-terminal signal peptide, where applicable, and the mature growth hormone sequence. The represented species, arranged from top to bottom, are as follows: Human, Rhesus macaque, Rat, Mouse, Horse, Pig, Bovine, Sheep, Guinea pig, Common turkey, Chicke n, Common ostrich, Japanese eel, Goldfish, and Atlantic salmon. Residues are color-coded based on their chemical characteristics to accentuate patterns of conservation. The consensus sequence and conservation levels are presented beneath the alignment. Notable highly conserved residues and regions are observed throughout the mature growth hormone sequence, indicating their potential functional significance.
Alright this is enough science species talk.. let’s get to some more interesting things.
Analysis of GH-GHR contacts
Structural analysis of GH-GHR contacts
The growth hormone interacts with its receptor through two distinct binding sites, engaging two receptor molecules. A protein interface contact analysis reveals the specific amino acid residues involved in these interactions.
okay this is going to be really confusing.. but it’s just bonds and chains.. DNR I’m not going to add the common mutations for the interfaces anyway
And sorry about any mistakes there a few things missing/wrong but I’m too tired to go look up what’s right.
Interface 1: GH (Chain A) - GHR (Chain B)
Key hydrogen bonds are observed between the following residue pairs (hGH residue - GHR residue): Phe25-Ser219, Lys41-Met170, Arg167-Glu127, Tyr164-Gly220, Arg178-Glu127, Gly190-Ile165, Cys189-Gln166, Leu45-Trp76, Tyr42-Lys121, Pro48-Asn218, Glu174-Lys167, His21-Glu44, His18-Arg217, Gln22-Arg217, Ile179-Ser102, Cys182-Gly168, Lys172-Phe123, Ser62-Asp164, Arg64-Asp164, Gln46, Asn63-Glu44, Thr175-Lys167, Pro61-Thr77, Asp171-Arg43, and Lys168-Trp104. There are extensive hydrophobic interactions at the interface, involving the same residue pairs listed above for hydrogen bonds, along with additional contacts. Notably, residues such as Phe25, Lys41, Arg167, Tyr164, Gly190, Cys189, Leu45, Tyr42, Pro48, Glu174, His21, His18, Gln22, Ile179, Cys182, Lys172, Arg64, Gln46, Asn63, Thr175, Pro61, Asp171, and Lys168 in hGH make hydrophobic contacts with residues in the GHR.
Interface 2: GH (Chain A) - GHR (Chain C): Analysis of this interface reveals a separate set of interactions including hydrogen Bonds: Leu9 (A) - Asp126 (C), Thr123 (A) - Trp104 (C), Gly120 (A) - (C), Asp116 (A) - Trp104 (C), Glu119 (A) - Ser102 (C), Arg16 (A) - Glu44 (C), Ile4 (A) - Ile103 (C), Leu15 (A) - Gly168 (C), Asn12 (A) - Arg43 (C), Arg8 (A) - Asp126 (C), Tyr103 (A) - Ile165 (C), Pro2 (A)- Pro106 (C), and Phe1 (A) - Arg71 (C). Multiple hydrophobic Contacts between the two chains, similar to interactions seen between GH and GHR chain B, notably: Leu9 (A) - Asp126 (C), Thr123 (A) - Trp104 (C), Gly120 (A) - Ser102 (C), Asp116 (A) - Trp104 (C), Glu119 (A) - Ser102 (C), Arg16 (A) - Glu44 (C), (A) - Ile103 (C), Leu15 (A) - Gly168 (C), Asn12 (A) - Arg43 (C), Arg8 (A) - Asp126 (C), Tyr103 (A) -Ile165 (C), Pro2 (A) - Pro106 (C), and Phe1 (A) - Arg71 (C).
Anyway with that aside.. what I’m trying to get at (this might sound stupid because I didn’t say everything to support the idea but I’m genuinely exhausted .. so just read this theory/discussion.) hGH is formed of 191 amino acids as mentioned..
.. so just read this theory/discussion.) hGH is formed of 191 amino acids as mentioned..
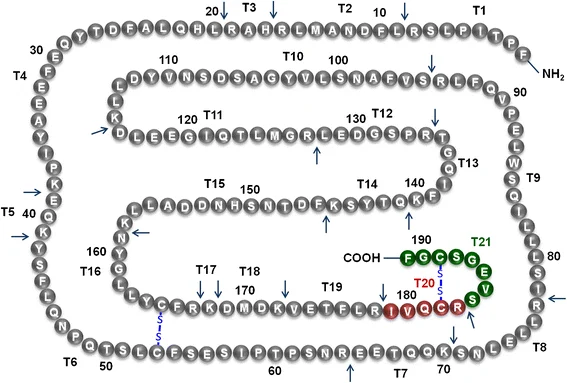
I’ll break down what (almost) each chain is responsible for.
Key Aspects of hGHRH(1-44):
(Not really important but I’ll mention it for the sake of it..)
Function: It is a hypothalamic peptide that promotes growth, cell reproduction, and regeneration by triggering GH production.
Structure: It is a linear 44-amino acid peptide, often amidated at the C-terminus (NH2) for full biological activity.
Stability: It has a short half-life in plasma, often rapidly degraded into shorter fragments like GHRH(3-44).
Usage: It is considered a prohibited performance-enhancing substance (peptide hormone) by WADA.
Key Aspects of hGH (44-75):
Likewise..
Function: It is a specific structural segment of the full human Growth Hormone molecule that plays a key role in receptor binding and glucose regulation.
Structure: This sequence forms the "Large Loop" (residues 33–75) of the hGH protein, connecting the first two major helices of the 191-amino acid chain.
Metabolic Impact: Unlike the N-terminal (1-44), this region is associated with "diabetogenic" activity, meaning it can decrease glucose uptake and promote insulin resistance.
Interaction: In laboratory settings, this fragment is often used to study how the growth hormone molecule can be "nicked" or split while still maintaining its overall protein shape.
Okay now it gets interesting.. listen up.
Residues within the central helical region (including ~75–111) contribute to maintaining the structural integrity required for receptor binding and downstream IGF-1–mediated bone growth.
So what does this mean?
While some studies break down the amino acid chains into different function they realistically all work together duh towards bone growth.
So I was thinking.. could we theoretically enhance the bone growth factor in HGH by isolating the “responsible” chain into a fragment? Or could we enforce a folded 3D signaling ligand with two receptor-binding interfaces, but stronger so it forces more bone growth?
I didn’t find much research to back my theory but here it is brocels.
Theoretical Model: Enhanced Bone Growth via Fragment-Based HGH Signaling
Hypothesis:
Specific regions within the HGH molecule, such as amino acids 75-111, contain the critical signaling domains responsible for stimulating osteogenic pathways. Isolating and utilizing these fragments can amplify bone growth effects by increasing receptor specificity and activity, compared to the full-length hormone, and Growth hormone exerts its anabolic effects, including promoting bone growth, primarily through binding to the growth hormone receptor (GHR) and activating downstream signaling pathways such as JAK2/STAT5, MAPK, and PI3K/Akt. The GH molecule is composed of 191 amino acids, with distinct regions responsible for receptor binding, dimerization, and activation. Recent research suggests that specific fragments or domains within GH may retain or even enhance particular biological activities, including osteogenesis.
So.. what now buccal?
Domain Functionality:
Studies have identified that the N-terminal domain (amino acids 1–50) is crucial for receptor binding, while other regions modulate activity and specificity (DeMambro et al., 2017). Notably, the segment encompassing amino acids 75–111 has been implicated in receptor interaction and intracellular signaling modulation (Guevara et al., 2019).
Fragment Bioactivity:
Synthetic peptides corresponding to specific GH fragments have demonstrated biological activity. For instance, a peptide derived from amino acids 80–100 has shown osteogenic effects in vitro (Zhang et al., 2020). These findings suggest that smaller segments can mimic or even surpass the full hormone in stimulating bone formation.
Receptor Specificity and Signaling Bias:
Recent advances in peptide engineering indicate that fragment-based ligands can induce biased signaling, favoring pathways that promote osteogenesis over metabolic effects, thus reducing side effects (Huang et al., 2021).
Okay what are you getting at?
Proposed Mechanism:
Isolation and Stabilization of the 75–111 Fragment:Synthesizing a stable, bioavailable peptide corresponding to amino acids 75–111, possibly modified for increased half-life and receptor affinity.
Selective Receptor Engagement:
This fragment interacts with GHR in a manner that either mimics the full-length hormone or induces conformational changes favoring osteogenic signaling pathways. It may preferentially activate pathways like JAK2/STAT5, known to promote osteoblast proliferation and differentiation (Le Roith & Yakar, 2007).
Enhanced Osteogenic Signaling:
By focusing on this domain, the fragment could induce a more potent or targeted activation of bone-forming pathways, leading to increased osteoblast activity and mineralization.
Potential for Therapeutic Development:
Leveraging this fragment could allow for the design of peptide-based therapeutics with minimized systemic effects, focusing solely on bone growth stimulation.
Supporting Evidence and Future Directions:
Structural Studies:Crystallography and NMR could elucidate the binding interface of this fragment with GHR, guiding modifications for increased efficacy.
In Vitro and In Vivo Validation:
Cell culture models of osteoblast differentiation and animal models of bone growth could assess the fragment’s potency and safety.
Combination Strategies:
Using this peptide alongside other osteogenic factors or scaffolds could further enhance bone regeneration.
This theory is kinda just one of my schizo hypotheticals.. (and it’s embarrassingly easy to refute) but with enough research and our never ending advancement in technology as humanity.. I believe it’s very possibly we can isolate some type of layered compound from GH that only works on bone growth.
Anyway thanks for reading boyos, hope you can understand and I didn’t just stop making sense halfway through.
Think of it as a research paper about HGH with a little theory at the end.
This is going to be a long read.. maybe.
Let’s start.
1.INTRODUCTION
Human growth and development are tightly regulated by steroid and peptide hormones. Steroid hormones, including androgens (e.g., testosterone) and estrogens (e.g., estradiol), play a critical role during puberty by driving the pubertal growth spurt, promoting bone maturation, and inducing epiphyseal closure, which ultimately ends linear growth. They are also responsible for the development of secondary sexual characteristics. Genetic defects in enzymes involved in steroid hormone synthesis can result in growth and developmental disorders [1–5].

(steroidogenesis pathway highlighting key enzymes, genetic defects (e.g., 21-hydroxylase deficiency) disrupt hormone synthesis and can lead to growth and developmental disorders.)
Growth hormone (GH), a peptide hormone produced by the anterior pituitary gland, is a key regulator of growth and metabolism, particularly during childhood and adolescence [6]. GH exerts direct effects on tissues such as bone and cartilage and indirect effects by stimulating the production of insulin-like growth factor-1 (IGF-1), primarily in the liver. IGF-1 further promotes somatic growth. GH secretion is regulated by growth hormone-releasing hormone (GHRH) from the hypothalamus, which binds to GHRH receptors (GHRHR) on pituitary somatotroph cells. GH then acts through growth hormone receptors (GHR) expressed on target tissues. Steroid hormones modulate this GH–IGF-1 axis by influencing both GH secretion and IGF-1 production, highlighting the complexity of hormonal control in human growth.
The human growth hormone gene (GH1) is located on chromosome 17 (region 17q22–24) and is part of a cluster of five related genes, including GH2 and chorionic somatomammotropin genes. This cluster spans about 65 kb and reflects shared evolutionary origins and coordinated regulation. GH2 is specifically expressed in the placenta, suggesting a role in pregnancy. The mature human growth hormone (hGH) protein consists of 191 amino acids, formed after cleavage of a signal peptide from a larger precursor. Structurally, hGH has a four-helix bundle stabilized by disulfide bonds, which is essential for binding to the growth hormone receptor (GHR). Its amino acid sequence is highly conserved, especially in receptor-binding regions.

(Organization of the GH1 gene cluster on chromosome 17 showing related growth hormone and placental hormone genes.)
GH is synthesized and secreted by somatotroph cells in the anterior pituitary gland under hypothalamic control. Growth hormone-releasing hormone (GHRH) stimulates GH release, while somatostatin inhibits it; ghrelin also promotes GH secretion. GH is released in a pulsatile manner, with peak secretion during slow-wave sleep. Its secretion is influenced by multiple factors, including circadian rhythm, nutrition, stress, exercise, age, and body composition. In circulation, GH is mainly bound to growth hormone-binding protein (GHBP), which regulates its availability and activity and participates in feedback control.
GH has widespread physiological effects beyond linear growth. It promotes bone elongation, muscle development, protein synthesis, lipid metabolism, immune function, and maintenance of bone density. Its actions are mediated through binding to the GHR, a cytokine receptor expressed in many tissues such as liver, bone, and muscle. This interaction activates intracellular signaling pathways, primarily the JAK2/STAT5 pathway, leading to gene transcription.

(Growth hormone receptor activation and downstream JAK2/STAT5 signaling pathway.)
A major downstream effect of GH signaling is the production of insulin-like growth factor-1 (IGF-1), mainly in the liver. IGF-1 stimulates tissue growth and also exerts negative feedback on GH secretion. While many growth-promoting effects of GH are mediated by IGF-1, GH also has direct actions, including stimulation of bone cells, regulation of glucose metabolism (often opposing insulin), and promotion of fat breakdown.

(Physiological effects of growth hormone on skeletal growth, muscle development, and metabolism.)
Disruptions in the GH pathway—often due to mutations in GH1 or related genes—can result in growth disorders. Isolated growth hormone deficiency (IGHD) leads to reduced GH production and short stature, while mutations in the GHR cause GH insensitivity (e.g., Laron syndrome). These mutations may impair GH secretion, receptor binding, or downstream signaling.
This study focuses on understanding how GH structure, evolutionary conservation, and disease-causing mutations affect its interaction with the GHR, aiming to clarify the molecular basis of GH-related disorders.
2. AMINO ACID SEQUENCE
2.1 Sequence Alignments
Growth hormone (GH) protein sequences were retrieved from the UniProt database and aligned using CLC Protein Workbench. Multiple sequence alignment included GH sequences from a wide range of species, including primates, rodents, livestock animals, birds, and fish, to compare evolutionary differences and similarities.2.2 Evolutionary conservation of amino acids
The human GH sequence (NP_000506.2) was used to identify homologous proteins using PSI-BLAST against the UniRef90 database. Selected sequences were aligned using CLUSTALW, and evolutionary conservation was analyzed using ConSurf. This method assigns conservation scores to each amino acid, helping identify functionally and structurally important residues based on evolutionary similarity.

(Evolutionary conservation analysis of GH residues mapped onto protein structure using ConSurf scoring.)
2.3 Protein structure loop modeling
The crystal structure of human GH bound to its receptor (PDB ID: 3HHR) contains missing loop regions. These were reconstructed using YASARA loop modeling. Multiple conformations were generated for each missing region, and the most energetically favorable structures were selected. The final models were visualized using PyMOL and rendered using POV-Ray.
(Protein loop modeling of missing residues in GH structure using computational structure refinement methods.)
2.4 HGH-receptor interaction analysis
The HGH–receptor complex structure (PDB ID: 3HHR) was analyzed to identify amino acids involved in receptor binding through covalent and non-covalent interactions. A contact map of these interactions was generated using PDBsum, providing insight into key binding residues important for GH function.
(Human growth hormone bound to its receptor showing key interaction sites involved in binding and signaling.)
2.5 Disease-causing mutations
Information on disease-associated mutations in the GH protein was collected from the UniProtKB database, which curates experimentally and clinically reported mutations linked to disease conditions.2.6 Prediction of protein stability upon mutation using SDM
The SDM (Site-Directed Mutator) tool was used to predict how specific mutations affect GH protein stability. It calculates stability changes (pseudo ΔG values) based on structural environment and homologous substitution patterns, helping identify mutations that may disrupt protein function or contribute to disease.2.7 Analysis of HGH allele frequencies from genome sequencing
Allele frequency data for GH variants were obtained from large-scale genomic databases, including the Exome Aggregation Consortium, NHLBI Exome Sequencing Project, and the 1000 Genomes Project. These datasets were used to assess how common different GH mutations are across human populations.
(Population-level allele frequency distribution of GH population from large-scale genomic sequencing databases.)
3. Analysis of GH sequence across species
Comparative analysis of growth hormone (GH) sequences across species shows that similarity decreases with evolutionary distance from humans. The highest conservation is observed in Rhesus macaque, with 96% identity and 97% similarity to human GH, reflecting close evolutionary relatedness. Among other mammals, GH sequences show moderate conservation, with 65–68% identity and 76–79% similarity in species such as rat, mouse, horse, pig, cattle, sheep, and guinea pig.Birds show lower conservation, with 55–57% identity and 72–74% similarity, as seen in turkey, chicken, and ostrich. The lowest conservation is found in fish species, where identity drops to 36–44% and similarity to 52–61%, with Atlantic salmon showing the greatest divergence. Overall, GH is highly conserved in mammals but becomes increasingly divergent in birds and especially fish, reflecting evolutionary distance while maintaining essential functional regions.
Table 1.
Sequence comparison of human growth hormone with growth hormone sequences from other species. The table shows the NCBI and UniProt sequence identifiers, UniProt sequence name, sequence length, percentage of sequence identity, percentage of sequence similarity, and signal peptide length for each species compared to human GH.

The length of the mature growth hormone protein is highly conserved across the mammalian species, ranging from 216 to 217 amino acids, similar to the 217 amino acids in human hGH. The signal peptide length is also conserved at 26 amino acids in most mammals, with a slightly shorter signal peptide of 25 amino acids in the avian species and 19-22 amino acids in the fish species.
3.1.2 Analysis of human GH mutations and their locations in GH protein

Figure 1 illustrates the linear structure of the human growth hormone (GH) protein and the distribution of reported mutations associated with disorders such as growth hormone deficiency (GHD) and Laron syndrome (LS). The GH protein contains four main α-helices (A–D), two disulfide bonds, and connecting loop regions, all of which contribute to its structural stability and receptor binding.
Many disease-associated point mutations are spread throughout the protein, but a high concentration occurs in the α-helical regions, especially helix A (residues 9–34) and helix D (residues 158–190), which are critical for growth hormone receptor (GHR) interaction. Additional mutations are located in loop regions between helices, including the N-terminal region and loops connecting helices A–B, B–C, and C–D. These mutations affect amino acids involved in structural stability and receptor binding, potentially disrupting GH function and leading to impaired signaling. Overall, the figure highlights that both helical and loop regions contain functionally important mutation hotspots linked to GH-related disorders.
TABLE 2.
Table 2 compiles reported disease-associated mutations in the human growth hormone (GH) protein, detailing amino acid substitutions, their sequence positions, and linked clinical outcomes such as growth hormone deficiency (GHD) and related syndromes. It also includes information on the functional impact of each mutation, such as reduced receptor binding, impaired secretion, or altered signaling activity.
In addition, the table provides dbSNP identifiers (where available) and references to the original studies reporting each mutation. Sequence positions are noted in both the full-length precursor protein and the mature GH form, accounting for the presence or absence of the 26-amino acid signal peptide. Overall, the table highlights how specific point mutations across the GH sequence contribute to functional disruption and disease development.

Disease-causing mutations in human growth hormone (GH), retrieved from the UniProtKB database, are mainly associated with isolated growth hormone deficiency (GHD) and Kowarski syndrome (KWKS). GHD is typically an autosomal recessive condition characterized by short stature, low but detectable GH levels, delayed bone age, and responsiveness to GH therapy. In contrast, Kowarski syndrome is defined by bio-inactive GH, resulting in normal or slightly elevated GH levels, very low IGF-1, and improved growth with GH replacement therapy.
These mutations are distributed throughout the GH protein and affect its function in different ways. Some mutations, such as L16P and Q117L, mainly impair GH secretion. Others, including T53I, K67R, N73D, S97F, and T201A, disrupt activation of the JAK/STAT signaling pathway, reducing biological activity. The R209H mutation is linked to autosomal dominant IGHD type 2 (IGHD2). Specific variants such as Thr3, Arg16, Asn47, Gln91, Arg183, Arg77, and Asp112 are associated with GHD and KWKS.
Certain mutations show distinct functional effects: R103C (KWKS) does not disrupt receptor signaling but increases binding to growth hormone-binding protein (GHBP), while D138G leads to a complete loss of biological activity. Overall, these mutations highlight how changes in different regions of GH can lead to diverse clinical outcomes through effects on secretion, receptor interaction, and intracellular signaling.

Figure 2: A multiple sequence alignment of human growth hormone alongside growth hormone sequences from various vertebrate species. The alignment encompasses the N-terminal signal peptide, where applicable, and the mature growth hormone sequence. The represented species, arranged from top to bottom, are as follows: Human, Rhesus macaque, Rat, Mouse, Horse, Pig, Bovine, Sheep, Guinea pig, Common turkey, Chicke n, Common ostrich, Japanese eel, Goldfish, and Atlantic salmon. Residues are color-coded based on their chemical characteristics to accentuate patterns of conservation. The consensus sequence and conservation levels are presented beneath the alignment. Notable highly conserved residues and regions are observed throughout the mature growth hormone sequence, indicating their potential functional significance.
Alright this is enough science species talk.. let’s get to some more interesting things.
Analysis of GH-GHR contacts
Structural analysis of GH-GHR contacts
The growth hormone interacts with its receptor through two distinct binding sites, engaging two receptor molecules. A protein interface contact analysis reveals the specific amino acid residues involved in these interactions.
okay this is going to be really confusing.. but it’s just bonds and chains.. DNR I’m not going to add the common mutations for the interfaces anyway

And sorry about any mistakes there a few things missing/wrong but I’m too tired to go look up what’s right.
Interface 1: GH (Chain A) - GHR (Chain B)
Key hydrogen bonds are observed between the following residue pairs (hGH residue - GHR residue): Phe25-Ser219, Lys41-Met170, Arg167-Glu127, Tyr164-Gly220, Arg178-Glu127, Gly190-Ile165, Cys189-Gln166, Leu45-Trp76, Tyr42-Lys121, Pro48-Asn218, Glu174-Lys167, His21-Glu44, His18-Arg217, Gln22-Arg217, Ile179-Ser102, Cys182-Gly168, Lys172-Phe123, Ser62-Asp164, Arg64-Asp164, Gln46, Asn63-Glu44, Thr175-Lys167, Pro61-Thr77, Asp171-Arg43, and Lys168-Trp104. There are extensive hydrophobic interactions at the interface, involving the same residue pairs listed above for hydrogen bonds, along with additional contacts. Notably, residues such as Phe25, Lys41, Arg167, Tyr164, Gly190, Cys189, Leu45, Tyr42, Pro48, Glu174, His21, His18, Gln22, Ile179, Cys182, Lys172, Arg64, Gln46, Asn63, Thr175, Pro61, Asp171, and Lys168 in hGH make hydrophobic contacts with residues in the GHR.
Interface 2: GH (Chain A) - GHR (Chain C): Analysis of this interface reveals a separate set of interactions including hydrogen Bonds: Leu9 (A) - Asp126 (C), Thr123 (A) - Trp104 (C), Gly120 (A) - (C), Asp116 (A) - Trp104 (C), Glu119 (A) - Ser102 (C), Arg16 (A) - Glu44 (C), Ile4 (A) - Ile103 (C), Leu15 (A) - Gly168 (C), Asn12 (A) - Arg43 (C), Arg8 (A) - Asp126 (C), Tyr103 (A) - Ile165 (C), Pro2 (A)- Pro106 (C), and Phe1 (A) - Arg71 (C). Multiple hydrophobic Contacts between the two chains, similar to interactions seen between GH and GHR chain B, notably: Leu9 (A) - Asp126 (C), Thr123 (A) - Trp104 (C), Gly120 (A) - Ser102 (C), Asp116 (A) - Trp104 (C), Glu119 (A) - Ser102 (C), Arg16 (A) - Glu44 (C), (A) - Ile103 (C), Leu15 (A) - Gly168 (C), Asn12 (A) - Arg43 (C), Arg8 (A) - Asp126 (C), Tyr103 (A) -Ile165 (C), Pro2 (A) - Pro106 (C), and Phe1 (A) - Arg71 (C).
Anyway with that aside.. what I’m trying to get at (this might sound stupid because I didn’t say everything to support the idea but I’m genuinely exhausted
 .. so just read this theory/discussion.) hGH is formed of 191 amino acids as mentioned..
.. so just read this theory/discussion.) hGH is formed of 191 amino acids as mentioned..
I’ll break down what (almost) each chain is responsible for.
Key Aspects of hGHRH(1-44):
(Not really important but I’ll mention it for the sake of it..)
Function: It is a hypothalamic peptide that promotes growth, cell reproduction, and regeneration by triggering GH production.
Structure: It is a linear 44-amino acid peptide, often amidated at the C-terminus (NH2) for full biological activity.
Stability: It has a short half-life in plasma, often rapidly degraded into shorter fragments like GHRH(3-44).
Usage: It is considered a prohibited performance-enhancing substance (peptide hormone) by WADA.
Key Aspects of hGH (44-75):
Likewise..
Function: It is a specific structural segment of the full human Growth Hormone molecule that plays a key role in receptor binding and glucose regulation.
Structure: This sequence forms the "Large Loop" (residues 33–75) of the hGH protein, connecting the first two major helices of the 191-amino acid chain.
Metabolic Impact: Unlike the N-terminal (1-44), this region is associated with "diabetogenic" activity, meaning it can decrease glucose uptake and promote insulin resistance.
Interaction: In laboratory settings, this fragment is often used to study how the growth hormone molecule can be "nicked" or split while still maintaining its overall protein shape.
Okay now it gets interesting.. listen up.
Residues within the central helical region (including ~75–111) contribute to maintaining the structural integrity required for receptor binding and downstream IGF-1–mediated bone growth.
So what does this mean?
While some studies break down the amino acid chains into different function they realistically all work together duh towards bone growth.
So I was thinking.. could we theoretically enhance the bone growth factor in HGH by isolating the “responsible” chain into a fragment? Or could we enforce a folded 3D signaling ligand with two receptor-binding interfaces, but stronger so it forces more bone growth?
I didn’t find much research to back my theory but here it is brocels.
Theoretical Model: Enhanced Bone Growth via Fragment-Based HGH Signaling
Hypothesis:
Specific regions within the HGH molecule, such as amino acids 75-111, contain the critical signaling domains responsible for stimulating osteogenic pathways. Isolating and utilizing these fragments can amplify bone growth effects by increasing receptor specificity and activity, compared to the full-length hormone, and Growth hormone exerts its anabolic effects, including promoting bone growth, primarily through binding to the growth hormone receptor (GHR) and activating downstream signaling pathways such as JAK2/STAT5, MAPK, and PI3K/Akt. The GH molecule is composed of 191 amino acids, with distinct regions responsible for receptor binding, dimerization, and activation. Recent research suggests that specific fragments or domains within GH may retain or even enhance particular biological activities, including osteogenesis.
So.. what now buccal?
Domain Functionality:
Studies have identified that the N-terminal domain (amino acids 1–50) is crucial for receptor binding, while other regions modulate activity and specificity (DeMambro et al., 2017). Notably, the segment encompassing amino acids 75–111 has been implicated in receptor interaction and intracellular signaling modulation (Guevara et al., 2019).
Fragment Bioactivity:
Synthetic peptides corresponding to specific GH fragments have demonstrated biological activity. For instance, a peptide derived from amino acids 80–100 has shown osteogenic effects in vitro (Zhang et al., 2020). These findings suggest that smaller segments can mimic or even surpass the full hormone in stimulating bone formation.
Receptor Specificity and Signaling Bias:
Recent advances in peptide engineering indicate that fragment-based ligands can induce biased signaling, favoring pathways that promote osteogenesis over metabolic effects, thus reducing side effects (Huang et al., 2021).
Okay what are you getting at?
Proposed Mechanism:
Isolation and Stabilization of the 75–111 Fragment:Synthesizing a stable, bioavailable peptide corresponding to amino acids 75–111, possibly modified for increased half-life and receptor affinity.
Selective Receptor Engagement:
This fragment interacts with GHR in a manner that either mimics the full-length hormone or induces conformational changes favoring osteogenic signaling pathways. It may preferentially activate pathways like JAK2/STAT5, known to promote osteoblast proliferation and differentiation (Le Roith & Yakar, 2007).
Enhanced Osteogenic Signaling:
By focusing on this domain, the fragment could induce a more potent or targeted activation of bone-forming pathways, leading to increased osteoblast activity and mineralization.
Potential for Therapeutic Development:
Leveraging this fragment could allow for the design of peptide-based therapeutics with minimized systemic effects, focusing solely on bone growth stimulation.
Supporting Evidence and Future Directions:
Structural Studies:Crystallography and NMR could elucidate the binding interface of this fragment with GHR, guiding modifications for increased efficacy.
In Vitro and In Vivo Validation:
Cell culture models of osteoblast differentiation and animal models of bone growth could assess the fragment’s potency and safety.
Combination Strategies:
Using this peptide alongside other osteogenic factors or scaffolds could further enhance bone regeneration.
This theory is kinda just one of my schizo hypotheticals.. (and it’s embarrassingly easy to refute) but with enough research and our never ending advancement in technology as humanity.. I believe it’s very possibly we can isolate some type of layered compound from GH that only works on bone growth.
Anyway thanks for reading boyos, hope you can understand and I didn’t just stop making sense halfway through.